 CoDockPP
CoDockPP
A multistage protein-protein docking program based on shape complementarity, knowledge-based scoring function and site constraint.
Submit | Check Result | Job Status | Result Example | Tutorial | References
 CoDockPP
CoDockPP

A multistage protein-protein docking program based on shape complementarity, knowledge-based scoring function and site constraint. Submit | Check Result | Job Status | Result Example | Tutorial | References |
You need to upload the input coordinates of both receptor (larger) and ligand (smaller) proteins in strict PDB format. The PDB checker, such as Molprobity, should be used to anticipate and fix any potential PDB error before you upload the pdb files. You should introduce an email address to receive a direct link to access your docking results later.
The codockpp can performs global docking and site-specific docking to predict the binding complexes between two proteins. You can enter one constraint residue on the receptor interface and another one on the ligand interface.
The binding site residues are provided in the text box as follows
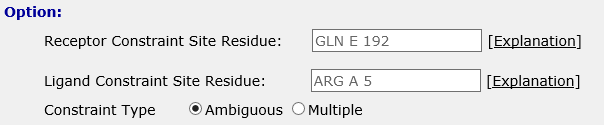
which stand for residue GLN192 of chain E on the receptor, and ARG5 of chain A on the ligand.
The site constraints can be set as ambiguous constraint and multiple constraint. When you choose ambiguous constraint, the conformation is required with at least one site on the interface of receptor or ligand. When you choose multiple constraint, the conformation is retained with both of the two sites on the interface.
After your job completes, entering your job ID will bring you to the results for that job.
Alternatively, you can access the docking results by the corresponding link in your email. The download url is provided as: http://codockpp.schanglab.org.cn/download.php?dir=*************
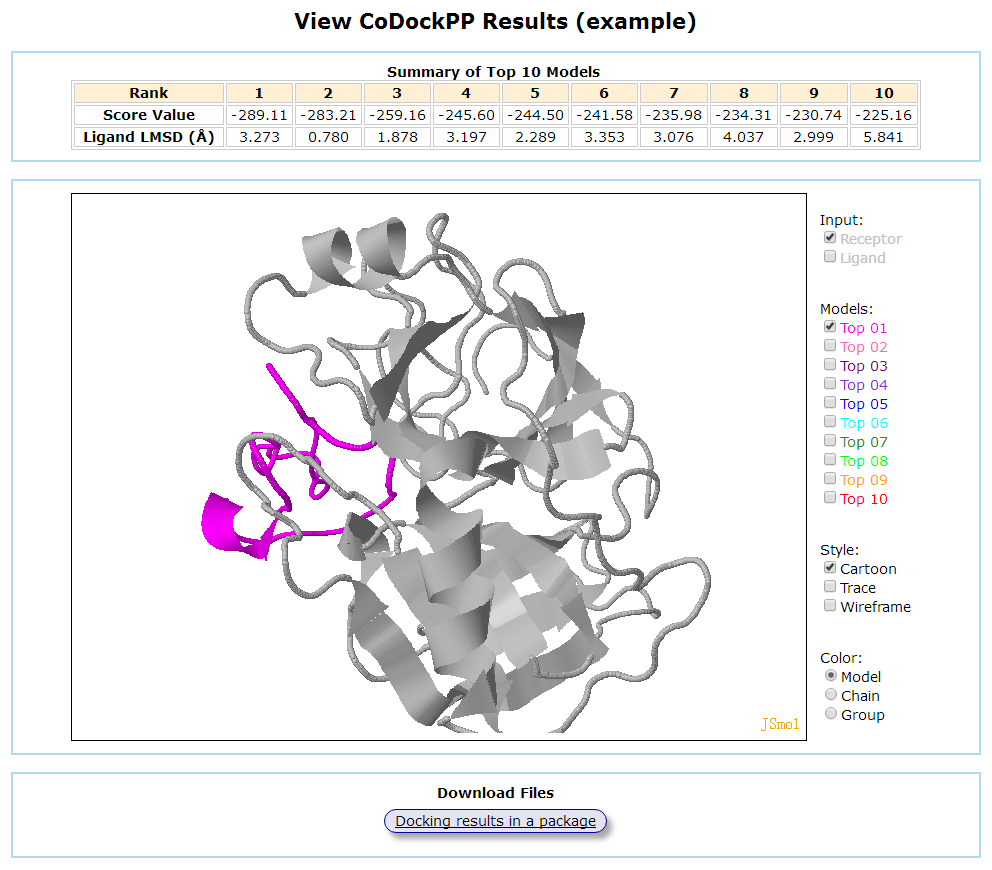
By default, the server only displays the top 10 models. The uploaded receptor is renamed Input_R.pdb, and the ligand is renamed Input_L.pdb. The top 10 models are named Top1.pdb-Top10.pdb, respectively. The models can be visualized in 3D by JSmol. The scored file is named codockpp.dat. Then, you can view and download the results of your docking.
If you have any suggestions or questions, please Contact Us.